Helicube
Helicube is an integrated solution for managing and analyzing Next Generation Sequencing (NGS) data. Preinstalled with state-of-the-art software BALSA and database.bio, and common pipelines such as BWA+GATK and TopHat, Helicube supports heterogeneous workloads, ranging from data analysis to results visualization. Helicube can serve multiple users via web browser. Helicube is designed to be HIPAA compliant; it allows users to share projects and data using carefully designed privileges, while strictly protecting private data. Helicube can scale up to multiple units to support the workloads of larger institutes; workload scheduling and balancing is also automatic, giving users convenience and efficiency.
Fast and accurate genome analysis:
From raw reads to variants, BALSA takes a few hours for WGS, and ten plus minutes for WES.
Bioinformatics best practices:
Pre-installed pipelines include BALSA, BWA+GATK, TOPHAT, STAR and Cufflinks. Extendable with Helicube Python SDK.
Interactive variant analysis:
Variant interpretation and pathogenicity classification with over 30 databases including ClinVar, TCGA, ICGC, EVS and dbSNP.
Easy sharing: Fine-grained privileges enabling contributors, analyzers, methodologists, etc. to collaborate on your project
Helicube features two independent systems: the Helicube Engine for genome analysis and database.bio for visualization. The two systems can be used separately, or as an integrated unit.
The Helicube Engine stores and manages user data and analysis results securely. It enables management of large genomics projects through fine-grained access settings for collaborators. Top-notch tools and best practices for NGS analysis including BALSA, BWA+GATK, TOPHAT, STAR and Cufflinks are pre-installed. As an advanced feature, Helicube provides an SDK for users to upload their own tools and create new pipelines.
database.bio is a clinical decision-support system that interprets genome variants (input with VCF and 23andMe formats). database.bio integrates genomic, clinical and pharmaceutical information from different sources into a uniform database in which variants are described with standard nomenclatures.
BALSA is an ultra-fast and accurate software for genome secondary analysis. Accelerated by a GPU, it is 5 or more times faster than BWA+GATK and can finish a WES analysis in minutes. It enables the application of NGS in the clinical context, where fast turn around time is required.
Benchmarking on the quality of variants using NIST’s “Genome in a Bottle” demonstrates that the variants called by BALSA has the highest combined sensitivity and precision (refer to Figure 2).
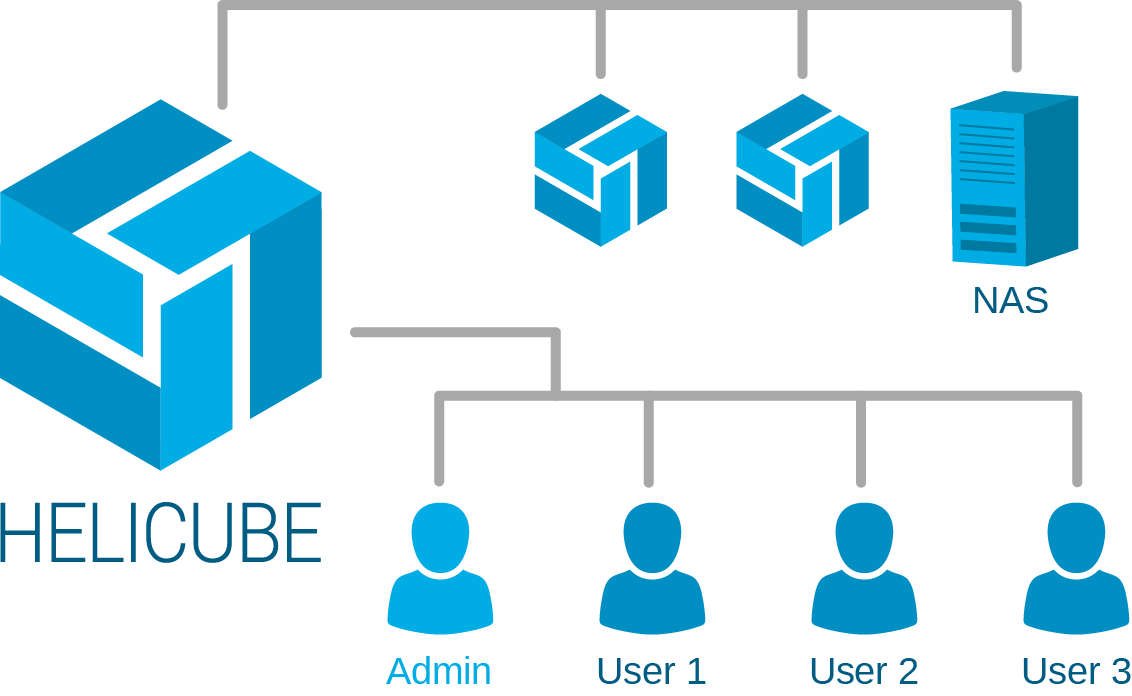
Helicube can scale up to more than ten hardware units to support the extensive workload of your institute. These units work coherently in a management free and efficient manner, enabling you to focus on your research instead of the infrastructure.
Helicube is designed to be HIPAA compliant and uses encryption standard AES256 for secure data transfer.
Helicube only runs authorized tools, ensuring the integrity of the data handled by the system. It also provides a monitoring function to give users timely updates about system health.